MEME
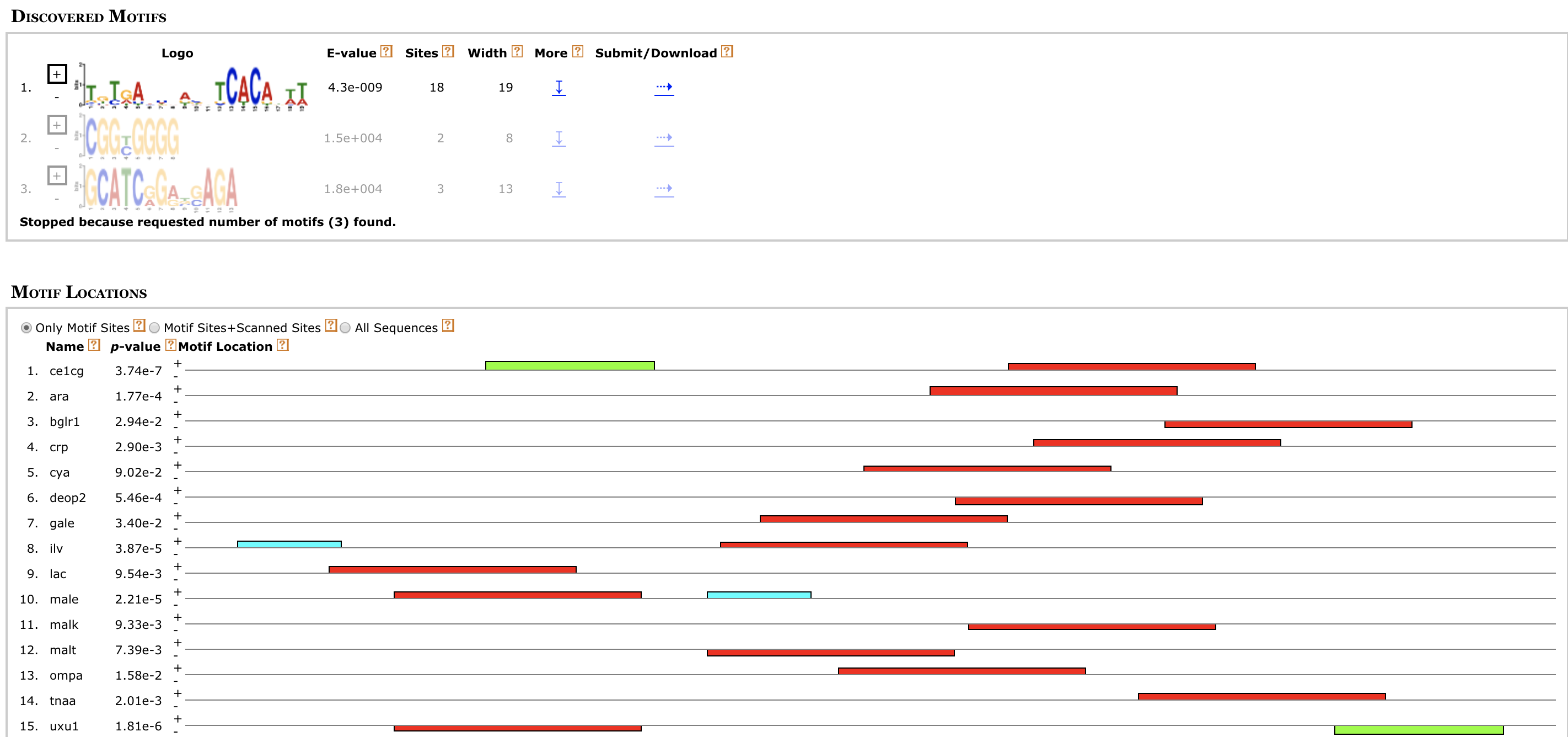 http://meme-suite.org/doc/examples/meme_example_output_files/meme.html?man_type=web
http://meme-suite.org/doc/examples/meme_example_output_files/meme.html?man_type=web
Methods
linear multiple view multiple scale single focus contiguous no abstraction no arrangement no interconnection point sparse type point contiguous typeTool
| Access Format | web application standalone app |
| Supported Files | fasta |
| License | The Regents of the University of California |
| Tool name | MEME |
| Tool Link | http://meme-suite.org/ |
| Documentation | http://meme-suite.org/doc/overview.html?man_type=web |
Paper
MEME: discovering and analyzing DNA and protein sequence motifs.
Bailey TL, Williams N, Misleh C, Li WW. MEME: discovering and analyzing DNA and protein sequence motifs. Nucleic Acids Res. academic.oup.com; 2006;34: W369–73.
Abstract
MEME (Multiple EM for Motif Elicitation) is one of the most widely used tools for searching for novel ‘signals’ in sets of biological sequences. Applications include the discovery of new transcription factor binding sites and protein domains. MEME works by searching for repeated, ungapped sequence patterns that occur in the DNA or protein sequences provided by the user. Users can perform MEME searches via the web server hosted by the National Biomedical Computation Resource ( http://meme.nbcr.net ) and several mirror sites. Through the same web server, users can also access the Motif Alignment and Search Tool to search sequence databases for matches to motifs encoded in several popular formats. By clicking on buttons in the MEME output, users can compare the motifs discovered in their input sequences with databases of known motifs, search sequence databases for matches to the motifs and display the motifs in various formats. This article describes the freely accessible web server and its architecture, and discusses ways to use MEME effectively to find new sequence patterns in biological sequences and analyze their significance.