CGView Server
 http://stothard.afns.ualberta.ca/cgview_server/examples/example_1/images/settings_1.png
http://stothard.afns.ualberta.ca/cgview_server/examples/example_1/images/settings_1.png
Methods
circular single view single scale single focus segregated no abstraction no arrangement no interconnection segment sparse type segment contiguous type point sparse type point contiguous typeTool
| Access Format | web application |
| Supported Files | fasta other |
| License | unavailable |
| Tool name | CGView Server |
| Tool Link | http://stothard.afns.ualberta.ca/cgview_server/ |
| Documentation | n/a |
Paper
The CGView Server: a comparative genomics tool for circular genomes.
Grant JR, Stothard P. The CGView Server: a comparative genomics tool for circular genomes. Nucleic Acids Res. academic.oup.com; 2008;36: W181–4.
Abstract
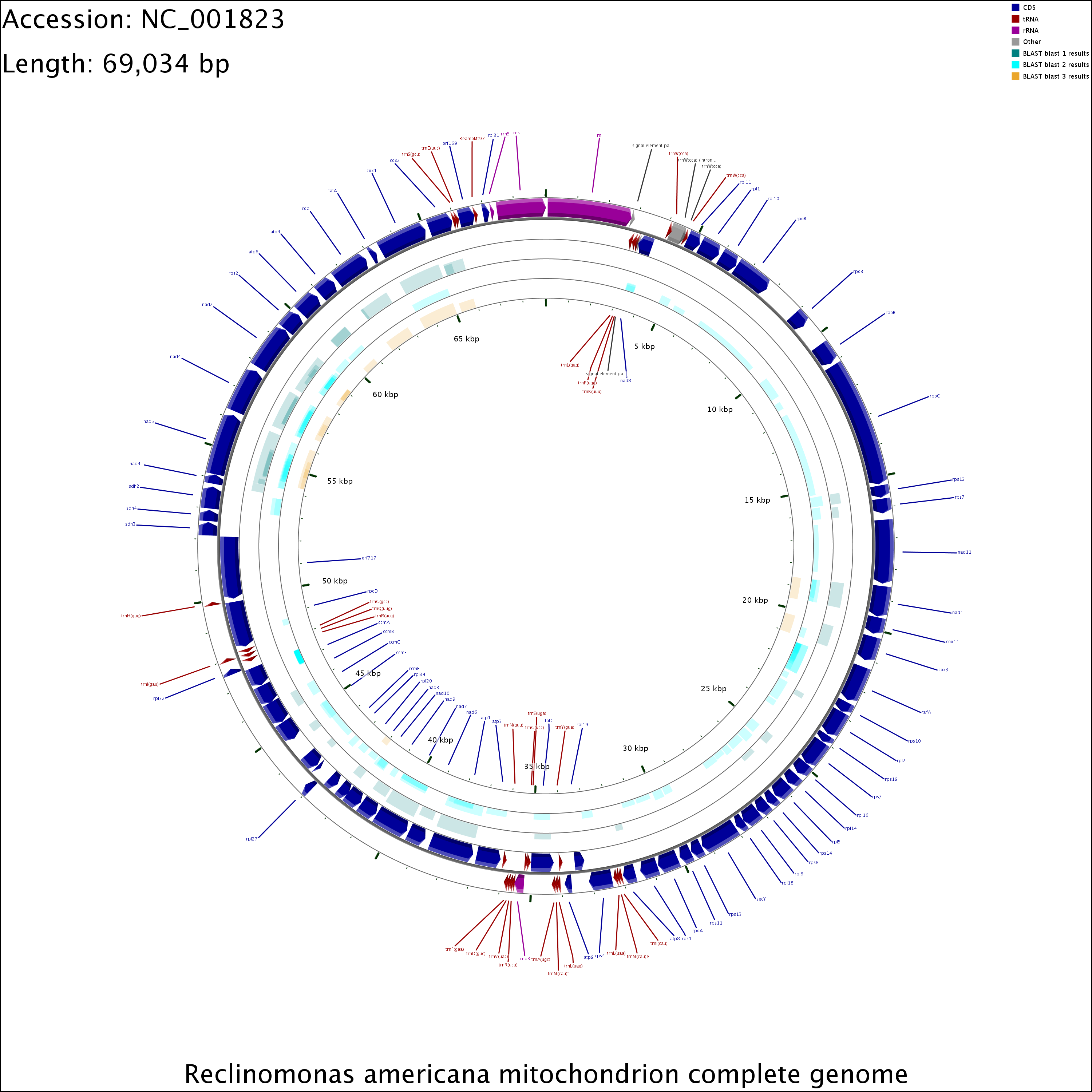
The CGView Server generates graphical maps of circular genomes that show sequence features, base composition plots, analysis results and sequence similarity plots. Sequences can be supplied in raw, FASTA, GenBank or EMBL format. Additional feature or analysis information can be submitted in the form of GFF (General Feature Format) files. The server uses BLAST to compare the primary sequence to up to three comparison genomes or sequence sets. The BLAST results and feature information are converted to a graphical map showing the entire sequence, or an expanded and more detailed view of a region of interest. Several options are included to control which types of features are displayed and how the features are drawn. The CGView Server can be used to visualize features associated with any bacterial, plasmid, chloroplast or mitochondrial genome, and can aid in the identification of conserved genome segments, instances of horizontal gene transfer, and differences in gene copy number. Because a collection of sequences can be used in place of a comparison genome, maps can also be used to visualize regions of a known genome covered by newly obtained sequence reads. The CGView Server can be accessed at http://stothard.afns.ualberta.ca/cgview_server/.