IRScope
 https://irscope.shinyapps.io/irapp/
https://irscope.shinyapps.io/irapp/
Methods
linear single view single scale single focus segregated partial abstraction no arrangement no interconnection segment sparse type segment contiguous typeTool
| Access Format | web application |
| Supported Files | fasta other |
| License | Under the license of the University of Helsinki |
| Tool name | IRScope |
| Tool Link | https://irscope.shinyapps.io/irapp/ |
| Documentation |
Paper
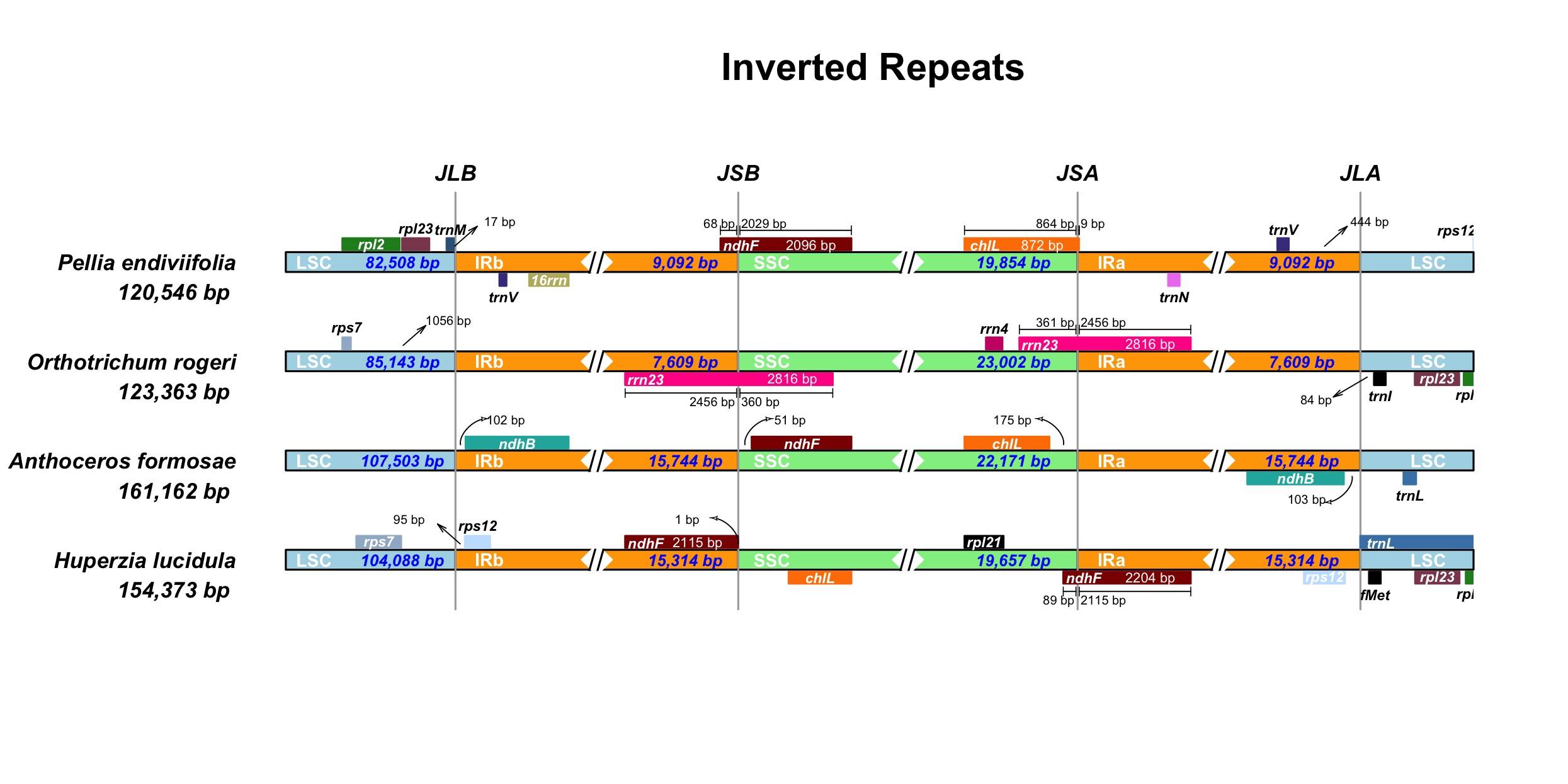
IRscope: an online program to visualize the junction sites of chloroplast genomes.
Amiryousefi A, Hyvönen J, Poczai P. IRscope: An online program to visualize the junction sites of chloroplast genomes. Bioinformatics. 2018; doi:10.1093/bioinformatics/bty220
Abstract
MOTIVATION: Genome plotting is performed using a wide range of visualizations tools each with emphasis on a different informative dimension of the genome. These tools can provide a deeper insight into the genomic structure of the organism. RESULTS: Here, we announce a new visualization tool that is specifically designed for chloroplast genomes. It allows the users to depict the genetic architecture of up to ten chloroplast genomes in the vicinity of the sites connecting the inverted repeats to the short and long single copy regions. The software and its dependent libraries are fully coded in R and the reflected plot is scaled up to realistic size of nucleotide base pairs in the vicinity of the junction sites. We introduce a website for easier use of the program and R source code of the software to be used in case of preferences to be changed and integrated into personal pipelines. The input of the program is an annotation GenBank (.gb) file, the accession or GI number of the sequence or a DOGMA output file. The software was tested using over a 100 embryophyte chloroplast genomes and in all cases a reliable output was obtained. AVAILABILITY AND IMPLEMENTATION: Source codes and the online suit available at https://irscope.shinyapps.io/irapp/ or https://github.com/Limpfrog/irscope.